labdata
labdata is a comprehensive software package designed to interact with databases and data file servers, enabling organized storage of complex datasets while facilitating reproducibility and efficient management of neuroscience research data.
This software is built on top of datajoint. Before proceeding, please familiarize yourself with datajoint's core concepts for building schemas and database querying. The system requires a database setup that can operate either locally or on AWS. For laboratory deployment, we recommend designating a dedicated administrator to manage users, AWS infrastructure, and access permissions.
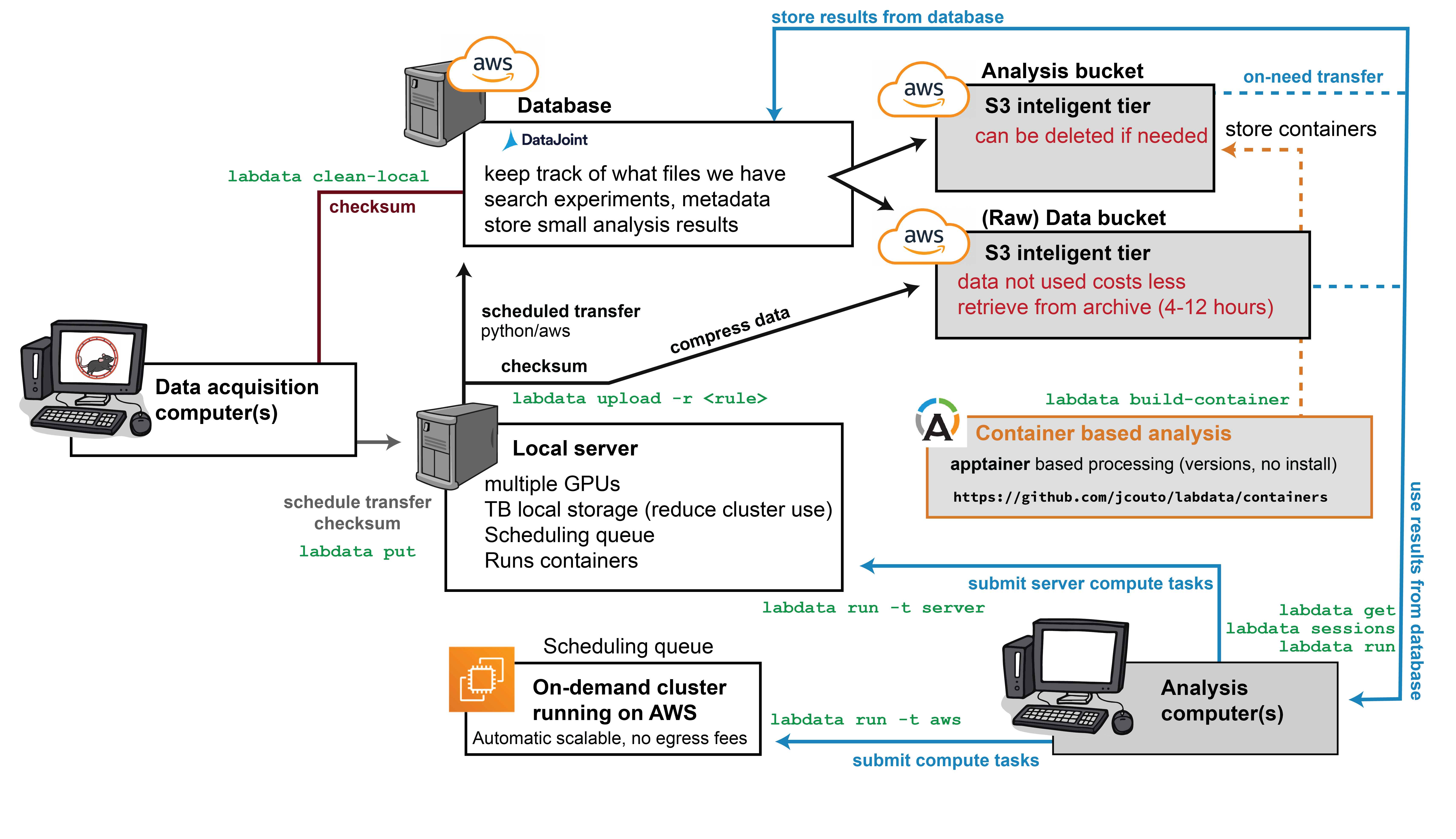
Overview
Experimental metadata and data file references are stored in a MySQL database, while actual data can be stored either on AWS or locally. This system is specifically designed to manage data and analysis workflows for medium-sized laboratories generating 100TB+ of data annually.
Core Features
labdata provides comprehensive functionality for:
- Database Interaction: Seamless interaction with the underlying database system
- Data Transfer Management: data transfer and cleanup from experimental computers
- Cloud Backup: File compression, checksum verification, and upload to cloud backup servers
- Data Retrieval: Efficient download and extraction of archived cloud data
- Stream Synchronization: Coordination of data from multiple experimental streams
- HPC Integration: Integration with high-performance computing clusters and cloud computing resources
- Containerized Analysis: Analysis execution using Apptainer containers
- Web-based Curation: Data curation and metadata insertion through a web browser interface
- Future Features:
- Database export functionality for collaborative sharing
- Automated archiving of legacy analysis results to reduce database size
Data Compression
The system implements data compression strategies (referred to as upload rules) for:
-
Imaging Datasets: Using zarr format with built-in support for:
- Widefield imaging
- Two-photon imaging
- Miniscope recordings
- Fixed whole-brain imaging
-
Electrophysiology Data: Using mtscomp for extracellular recordings
Analysis Capabilities
Data analysis workflows are implemented as compute tasks using Apptainer containers, providing extensible and reproducible analysis pipelines. The following analysis types are currently deployed and ready for use:
- Cell Segmentation:
- CaImAn for calcium imaging analysis
- Suite2p for two-photon imaging processing
- Spike Sorting: Multiple versions of Kilosort for neural spike detection and clustering
- Pose Estimation: DeepLabCut for behavioral tracking and pose analysis
For feedback and support, please contact: jcouto@gmail.com